Cancer Cytogenetics
Cytogenetics and cancer
Canine cancer
It is estimated that almost 50% of all dogs over the age of 10 years will develop cancer and approx 1 in 4 of all dogs will at some stage in their life develop a cancer. Not all cancers kill dogs and the high level of veterinary care that is available in the 21st century is providing man’s best friend with a very high quality of medical care.
Our research activities are focused on exploring the cellular genomics of canine cancers and the investigation of any evolutionarily related genetic aetiology shared with human.
Our lab is focused:
- the comparative aspects of genetic changes in canine and human cancers,
- the identification of cancer-related genes in dog and their counterparts in human cancer.
This work involves three major areas:
Direct molecular cytogenetics analysis of tumor cells. Tumor specific chromosome aberrations are widely documented in human malignancies and cytogenetics plays a key role in the diagnosis and prognosis of many forms of human cancer. To help with dog chromosome identification we have developed extensive panels of chromosome specific single locus probes (SLP) (numbering over 1,500 SLPs) and of whole chromosome paint probes (WCPP) that we use in multicolor fluorescence in situ hybridization (FISH) in order to characterize genetic aberrations in individual tumor cells. Use of a panel of 41 multicolored SLPs to identify each chromosome in the canine karyotype is shown below.
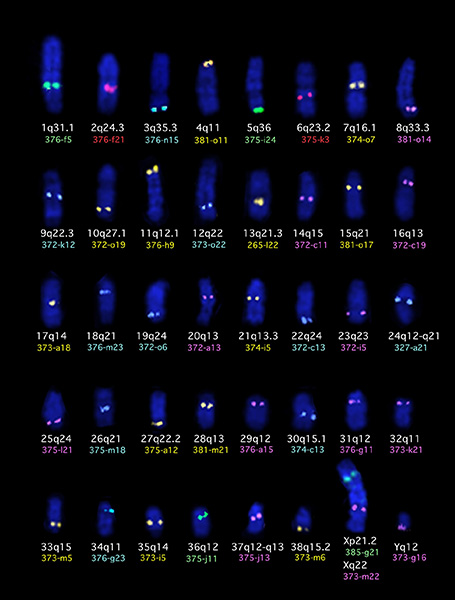
Indirect molecular cytogenetics analysis of tumor cells. Comparative genomic hybridization (CGH) is a whole genome screening approach that allows detection and localization of DNA sequence copy number variation anywhere in the tumor genome. The approach is based on competitive differential hybridization of labeled genomic DNA from a tumor, competed with DNA from normal cells. CGH has proved very helpful in defining the locations of previously unknown oncogenes and tumor suppressor genes. We have developed canine CGH and are now applying this technique to a variety of key tumor types. We have already begun to find associations between genetic changes detected by CGH analysis and the survival of dogs with certain cancers.
To improve the throughput, resolution and accessibility of canine CGH, we developed array-based CGH for the dog, first with a panel of 2,200 canine BAC clones ordered throughout the genome at 1Mb intervals, and now with a set of 1,000,000 oligonucleotide sequences spaced at 2.5kb intervals. The increase in resolution and throughput has led to array CGH replacing chromosome based CGH as the method of choice for genome wide assessment of DNA copy number aberrations.
The use of laser microdissection allows us to accurately micro-dissect tumor cells from archival sections. Using degenerate oligonucelotide primer (DOP) PCR, and fluorescent labeling, we are able to perform CGH using DNA obtained from just a few tumor cell nuclei. By characterizing the presence and location of chromosomal changes in the earliest identifiable changes in tumor cells, we are identifying possible targets for positional cloning and gene-based therapies.
Gene expression profiling. We are also developing array-based expression profiling to compare gene expression patterns in tumor cells compared to the pattern of expression of normal cells in adjacent tissue. A combination of the data obtained from genomic and expression arrays will allow us to 1) identity regions of the genome imbalance and then 2) define deregulated genes within these regions. In this way we are defining loci for mutation screening.